In the last six years, the cost of sequencing DNA – mapping out the genetic make-up of a given organism – has fallen staggeringly.
According to the US government's National Human Genome Research Institute, the cost of sequencing a million base pairs of DNA dropped from around $1,000 in 2006 to under 10 cents in 2012.
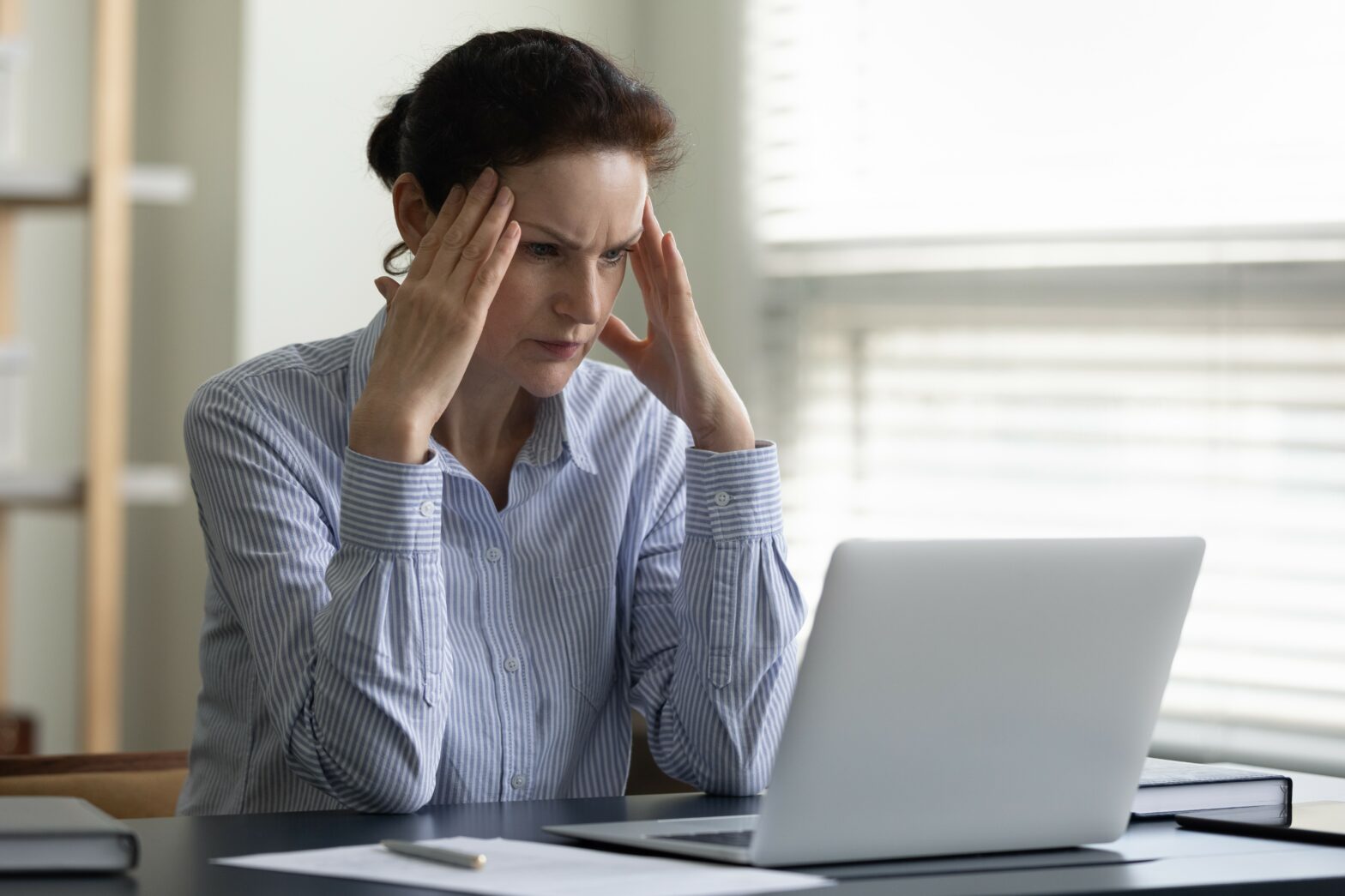
This is turning biology into a data-intensive science. The ability to sequence a specimen's DNA for as a little as a few dollars means researchers are generating reams and reams of genetic data, and, of course, using computational analysis to examine it.
This is just as true for commercial research and development (R&D) as it is for academic work. Anglo-Dutch consumer goods maker Unilever, for example, is pursuing a strategy it calls 'e-science'.
"The idea is to combine the data we generate in our R&D laboratories with publicly available biodata and analyse them together, taking an 'in silico' research approach wherever possible," explains Pete Keeley, technical IT lead for e-science at Unilever R&D.
This includes analysing microbial genetic data in order to understand how microbes interact with the human body so they can develop novel products.
"To make a better deodorant, for example, we might investigate the microbes that live in your armpits," explains Keeley. "If we can find which microbes are producing the body odour, and identify the active genes, we might be able to find a way to deactivate those genes or kill those microbes."
Microbial samples taken from various parts of the human body are sequenced by partner organisations and each sample can generate up to 100 giga bases of DNA.
This DNA data needs to be cleaned up and processed before it can be analysed. The sequences are small chunks of the genome, called 'short reads', that must be put back together to get the whole picture. Assembling millions of fragments of bacterial DNA from hundreds of species back into the correct order is like completing a huge, complex jigsaw puzzle.
Until 18 months ago, Unilever would process the genetic data using a grid cluster at its Port Sunlight laboratory in the Wirral. But the amount of data it needs to process will soon outstrip the capacity of that cluster, says Keeley.
"Every aspect of data sequencing is going to grow," he says. This includes both the number of samples Unilever expects to sequence and the resolution (and therefore file size) of each sequence.
"So the question was, how are we going to adapt? Are we going to have to double the size of our cluster every year or so, or is there a better way?
"Cloud," Keeley says, "was an obvious option to investigate."
Capacity planning
The appeal of cloud computing, he explains, was that it would allow Unilever to pay for the necessary compute capacity only when it was needed. "If we built a cluster that was big enough, we might only use it once a week," he says. "And we might only use it at 100% capacity once a year."
Unilever worked with Eagle Genomics, a consultancy that specialises in the application of cloud computing to 'bioinformatics', the computational analysis of large scale biological data.
Together, they developed a proof of concept implementation., Sequence data from microbial samples is uploaded to Amazon's S3 cloud storage service via the Internet. That data is then processed using specialist software such as gene sequence alignment program BLAST, deployed on EC2 instances.
When Unilever and Eagle Genomics started the project, Amazon’s cloud was the only real option, Keeley says. "They are quite a long way ahead of the competition in terms of working with scientific data."
The proof of concept was enough of a success that Unilever is now using Amazon Web Services in production.
As it turns out, the amount genetic data that Unilever needs to process has not grown as fast as its scientists were predicting 18 months ago. Still, it will happen inevitably, Keeley says, and in the mean time, cloud computing is a cost effective way to acquire compute capacity.
Unilever now plans to use the Amazon cloud to support genetic sequencing at other research labs around the world, including those in the US and China. That will happen within the year, Keeley says, although there are some legal issues to be considered first.